Based on these findings and the hypothesis that we proposed, new mutants by knocking off rif Q were developed with the expectation of increasing the production yield of the analog.
As stated above we proposed that in the mutant DCO34/36, 24-desmethylrifamycin B has a poor binding affinity to RifQ. RifQ so produced binds to RifP, thus, sufficient RifP is not available in DCO34/36 to transport 24-desmethyl rifamycin B. Based on these findings (Singhvi et al. 2020, 2021), a strategy was designed to produce new mutants by knocking off rifQ gene of rifamycin biosynthetic gene cluster in DCO 34/36 with the hope of increasing the production /yield of the analog significantly.
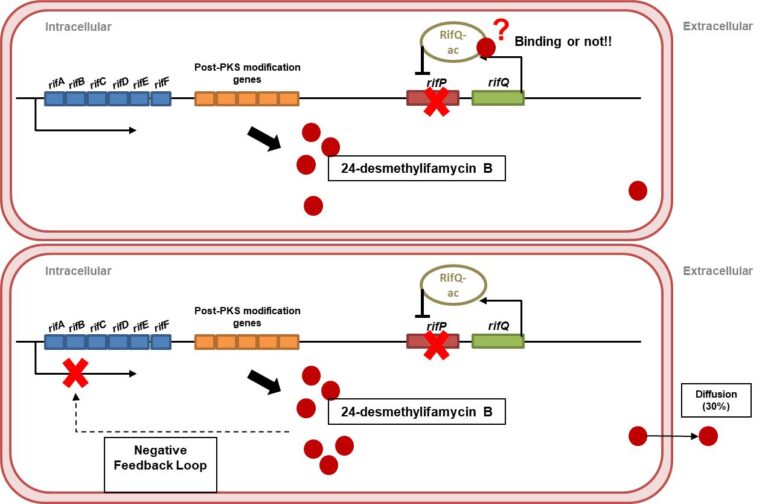
Figure 1: Mechanism of extracellular export of 24-desmethylrifamycin B in DCO34/36. depicting the role of rifQ and rifP in the transport of the molecule across the plasma membrane of A. mediterranei (Singhvi et al. 2021).
To validate this model, we thus went ahead to develop a method for creating knockout mutants of rifQ. The methodical procedure of homologous recombination by the use of a modified PCR-targeted Streptomyces gene replacement method to replace a gene of interest in Amycolatopsis mediterranei strains was developed. The knockout was carried out accordingly for rifQ in DCO36 and rifO in Amycolatopsis mediterranei S699. HR-homologous regions (2.5 kb) See Figure 6 (Müller et al. 2024).
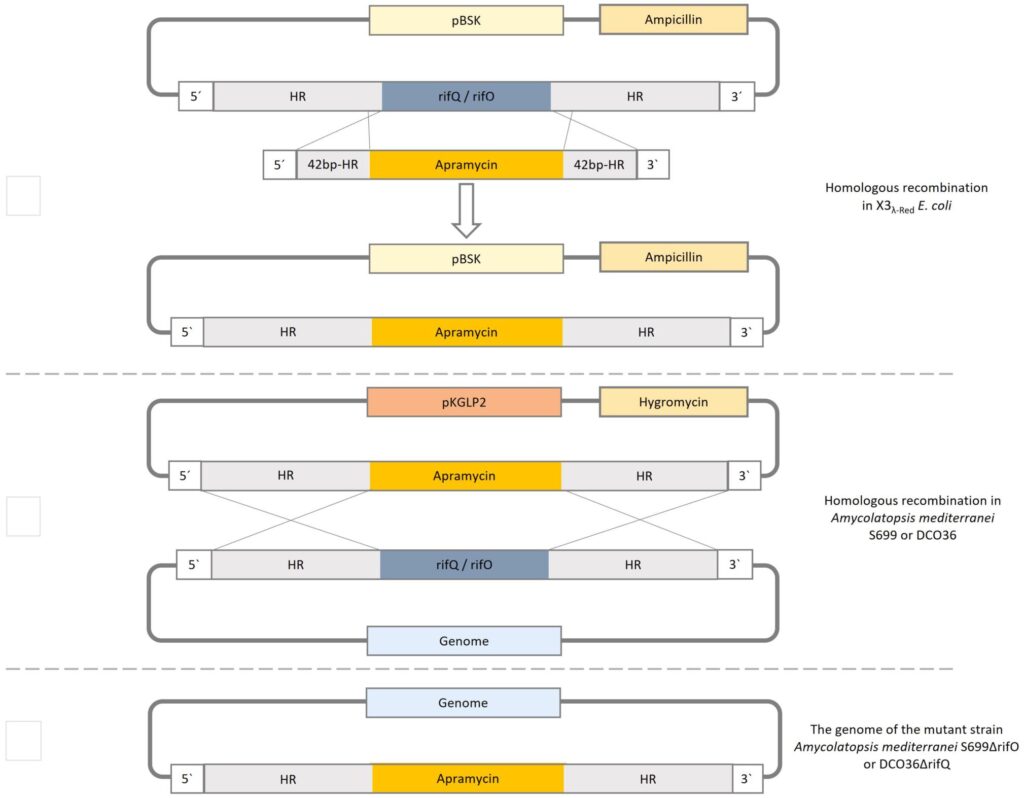
Figure 2: The methodical procedure of homologous recombination. (A) Homologous recombination in E coli X3l-Red cells to exchange the gene of interest (rifQ/rifO) and replace it with apramycin resistance cassette of pLERE to obtain pBSK-apraR-HR(rifQ/ rifO). (B) The construct apraR-HR(rifQ/ rifO) is integrated into the non-replicative suicidal Streptomyces vector pKLGP2 and transformed into ET12567/pUZ8002 for intergeneric conjugation with Amycolatopsis mediterranei S699 or DCO36. (C) The resulting genome after the replacement of the gene of interest (rifQ) with the apramycin resistance cassette (ApraR)(Müller et al.).
The mutant-generated produced has an increased production of 24-desmethylrifamycin B by 61.57% (Müller, M.; Bialas, E.; Sturm, I.; Sood, U.; Lal, R.; Bechthold, A, Frontiers in Antibiotics 2024, 3 https://doi.org/10.3389/frabi.2024.1399139).
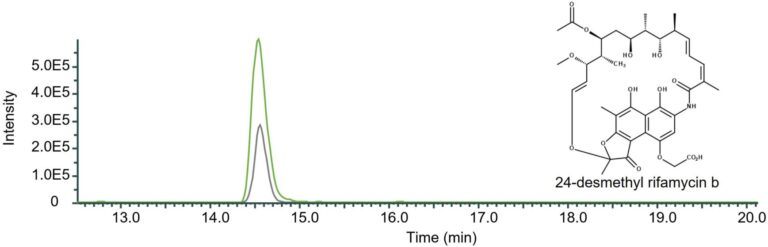
Figure 3: Qualitative comparison of the impact of the removal of rifQ in 24-desmethyl rifamycin B-producing mutant strain DCO36. The single mass-ion chromatogram shows the characteristic peak of the derivative 24-desmethyl rifamycin B m/z 740 [M-H] ─ at its retention time (Rt 14.50 min) resulting from HPLC-electrospray ionization mass spectrometry analysis of DCO36 (AUC = 2.722E6; grey) and DCO36ΔrifQ (AUC = 7.083E6; green) extracts. The comparison shows the increased amount of 24-desmethylrifamycin B after the removal of rifQ in a rifamycin B derivative-producing mutant strain by 61.57% (Mueller et al. 2024)
