Lorem ipsum dolor sit amet, consectetur adipiscing elit. Ut elit tellus, luctus nec ullamcorper mattis, pulvinar dapibus leo.
Project No: TE-16015/11/2019-TECH5 NMCG
Coordinating Institute
Partner Institutes:
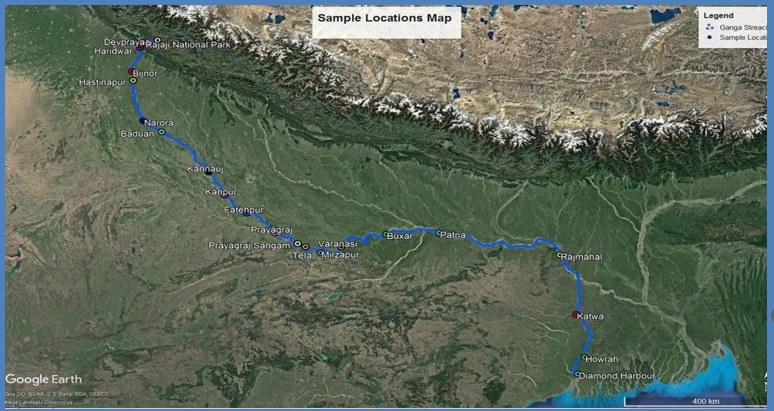
Microbes play a key role in regulating the ecosystem of any niche in the environment. Their adaptability to diverse conditions and their unique genotype, provide ecosystem services that contribute in maintaining a healthy niche and stability of an ecosystem. This project aims to understand the ecosystem services of microbes in a river niche, taking the river Ganga as a specific target. The Ganga river is a lifeline for those living along its path; it provides freshwater and is a water resource for agriculture, fisheries, industries, energy and tourism. Besides, this river has a deep social, cultural and religious footprint in India and houses a wide biodiversity. Microbes are one of the key drivers that maintain the health of the river. They help regulate the bio-geochemical cycles, carry out biodegradation of harmful organics (anthropogenic and natural), provide nutrients and growth factors to sustain aquatic life and are an indicator of pollution levels. Since 95% of the microbial world is unreported via culture-based studies, this project has used metagenomics to analyze microbes in the river Ganga based on their DNA.
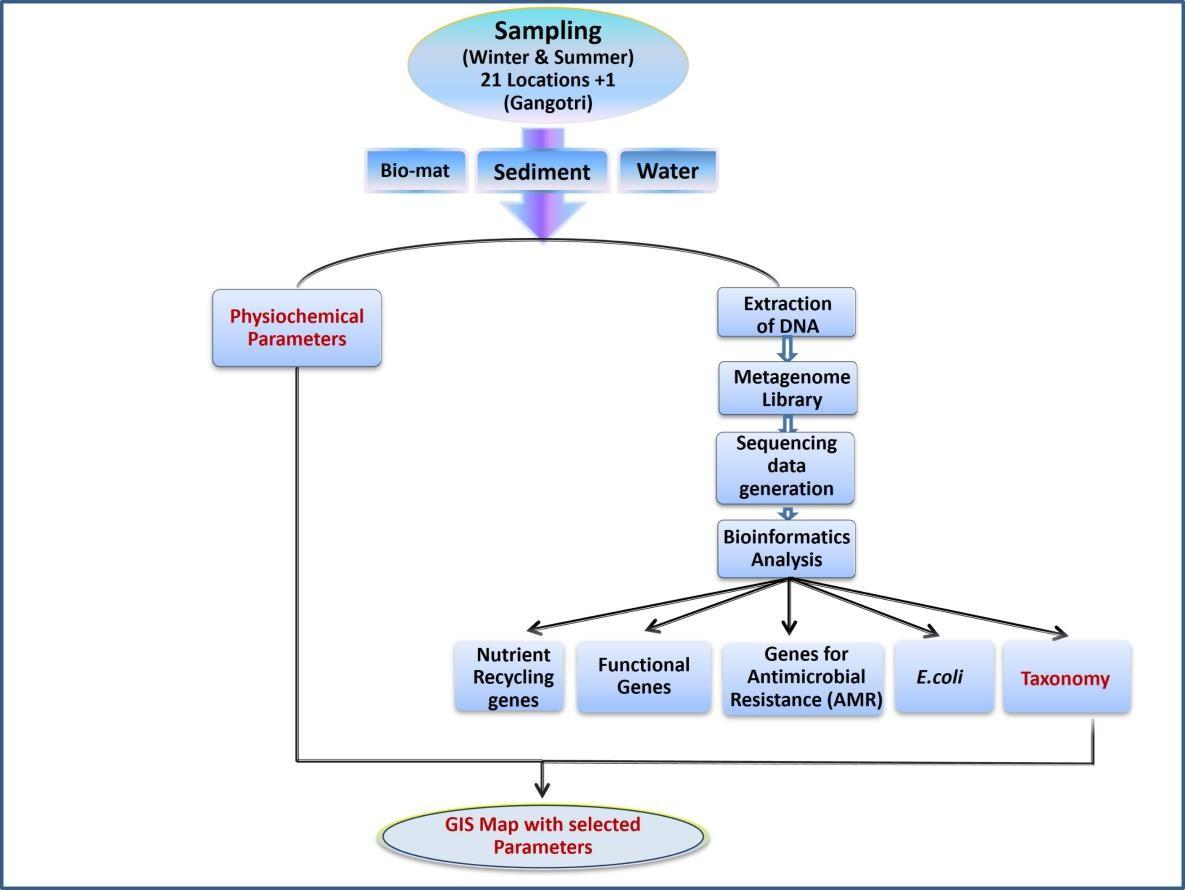
Figure 1: Distribution of the top ten genera in of the microbiome of River Ganga at different locations from Summer sampling time point
Figure 2: Heatmap showing the number of AMR associated genes determined in each sample for resistance to different classes of antibiotics. KA- Katwa, HW- Howrah, DH diamod Harbor. B stands Biomat, S- for Sediment and W for water samples.
