As mentioned above during subsequent research, after 2014, we identified bottlenecks in the production process of 24-desmethyl rifamycin B in the mutant strains. After generating the mutant DCO34/36 that produces 24 desmethyl rifamycin B that we published 10 years ago (Nigam et al., 2014), we again. checked for the efficacy of this drug. We found that the semisynthetic derivative of this analogue 24-desmethyl rifampicin as reported previously is nearly 50 times more effective than rifampicin against resistant strains of M. tuberculosis. However, there was a limitation of the low production of 24-desmethylrifamycin B in DCO34/36 mutants of A. mediterranei.
This was not unexpected as the yields of engineered strains through combinatorial approaches have been reported to be much lower than the parent compound during AT (acytransferase) swaps of the oleandomycin and in nystatin PKS swapping of a methyl-malonyl-CoA-specific AT domain (Long et al. 2002; Brautaset et al. 2003). While many other external and internal factors perhaps contribute to the low production of rifamycin B analogues in A. mediterranei S699, we focused after 2014 to understand better the regulation and export of rifamycin B and its analogues in the wild type and mutants so generated with a hope to overcome this limitation and thus improving the production of analogues in the mutants. Although it was a difficult job, however, finally we could resolve it (Müller, M.; Bialas, E.; Sturm, I.; Sood, U.; Lal, R.; Bechthold, A, Frontiers in Antibiotics 2024, 3. https://doi.org/10.3389/frabi.2024.1399139.
To resolve this, we first decided to carry out fermentation to increase the production, this however, did not solve the problem thus we focused on looking into the expression of structural and regulatory genes particularly of region IV of rifamycin biosynthetic gene cluster to study their expression during rifamycin production (Singhvi et al. 2021, Sood et al. 2024).
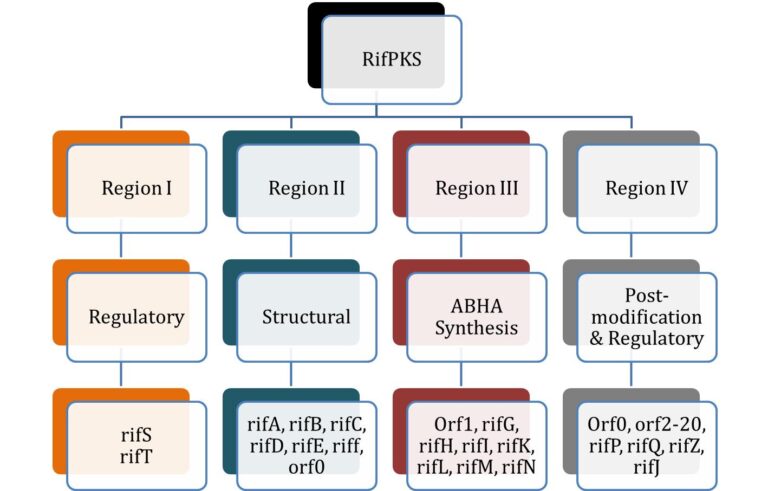
Figure 1: The genetic organization of the RifPKS (Rifamycin Polyketide Synthase) gene cluster, which is divided into four regions (Region I, Region II, Region III, Region IV). Each region contains specific genes categorized into various functional groups: regulatory, structural, ABHA synthesis, and post-modification & regulatory. (From Singhvi et al. 2021).
For this, we first conducted the proteomic analysis by selecting three strains of A. mediterranei that we had generated during our previous study (Wild type strain (S699 that produces rifamycin B), single cross-over (SCO) that did not produce rifamycin B) and the double cross over (that was generated in 2014 and produced 24-desmethyl rifamycin B -DCO34/36) . These strains provided good material to look more closely at the synthesis and regulatory pathways for the production of rifamycin B and 24 -desmethyl rifamycin Bi n A.mediterranei S699 and mutant DCO34/36 in particular. We envisaged that these findings will help us to further devise strategies to create mutants that can be used for better production of the analogue/s.
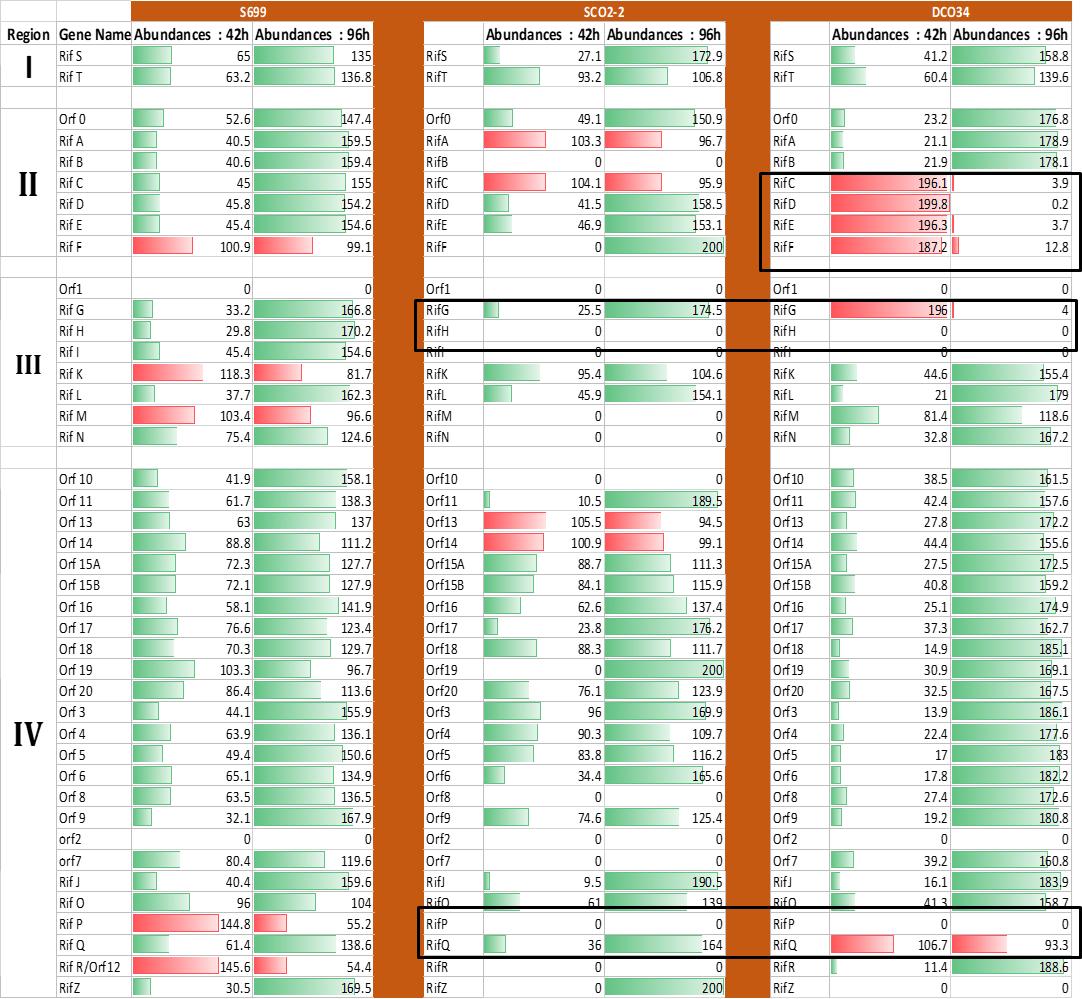
Figure 2: Comparative Expression profile of Amycolatopsis mediterranei S699, SCO2-2, and DCO34 rifamycin biosynthetic cluster at 42h and 96h. Change in expression is represented by bar where the green bar shows upregulation and the red bar denotes downregulation of genes from 42h to 96h (Singhvi et al. 2021).
Our studies on the proteomics analysis revealed that the structural genes (n=6) are downregulated in the mutant as a consequence of little or no expression of rifQ gene which is the key regulator and regulates rif P gene that in turn produce transmembrane proteins for the export of rifamycin B.
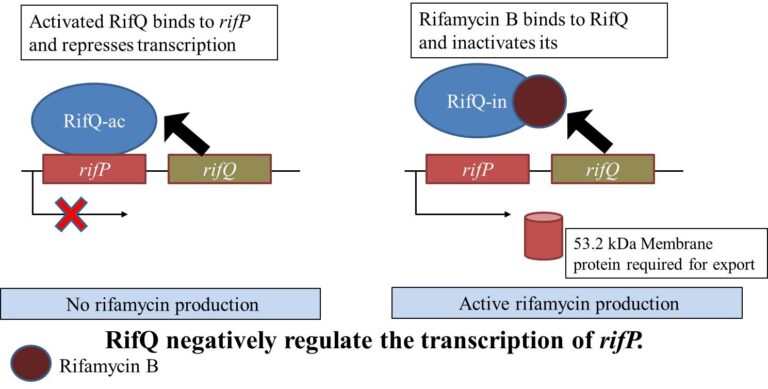
Figure 3: Role of rifQ in regulating the transcription of rifP. rifP is responsible for incorporating rifamycin transporters. rifQ negatively regulates the transcription of rifP, leading to lower transportation of rifamycin outside the cell. Rifamycin B binds and inactivates rifQ ( Lei et al. 2017 and Singhvi et al. 2021.)
Based on the earlier hypothesis by Lei et al. 2017, we proposed that rifQ is a key regulator for limiting the production of 24 desmethyl rifamycin B in DCO 34/36 (Singhvi et al., 2020; Singhvi et al. 2021). Thus, based on the previous work by Lei et al. 2017 we proposed a new hypothesis that the weak binding of 24 desmethyl rifamycin B with rifQ is responsible for the low production of 24-desmethyl rifamycin B in DCO 34/36 (Singhvi et al. 2021).
In brief, the data generated in these studies (Singhvi et al. 2020, 2021) and the information on the regulatory mechanism of rifamycin B already available in the literature (Li et al. 2017) reflected that the genes encoding RifQ and RifP are the main regulatory elements responsible for the alteration in the production of the analogue in DCO34/36. We thus proposed that 24-desmethyl rifamycin B either loosely binds to RifQ or not at all in in-turn, the rif P gene is not activated and transmembrane proteins are not produced (Singhvi et al. 2020, 2021). This prompted us to create rifQ mutants in DCO34/36.
